Transposable elements (TEs), which shift segments of the Genome are also known as “jumping genes”. These elements were first identified more than 50 years ago by geneticist Barbara McClintock of the Cold Spring Harbor Laboratory in New York. Biologists were initially sceptical of McClintock’s discovery. However, over the next several decades, it became clear that TEs not only “jump” but are also found in almost all organisms (both prokaryotes and eukaryotes), and usually in large numbers. For example, TEs constitute approximately 50% of the human genome and up to 90% of the maize genome (SanMiguel, 1996).
Retrotransposons
Unlike class 2 elements, class 1 elements, also known as retrotransposons, move through the action of RNA intermediates. In other words, class 1 TEs do not encode transposase; rather, they produce RNA transcripts and then rely on reverse transcriptase enzymes to reverse transcribe the RNA sequences back into DNA, which is then inserted into the target site.
There are two main types of class 1 TEs: LTR retrotransposons, which are characterized by the presence of long terminal repeats (LTRs) at both ends; and non-LTR TE, which lacks repeats. Both the LINE1, or L1, and Alu genes represent non-LTR TE families. L1 elements average about 6 kilobases in length. By contrast, Alu elements average only a few hundred nucleotides, making them a short interspersed transposable element, or SINE.
Alu is particularly prolific, originating in primates and expanding in a relatively short time to about 1 million copies per cell in humans. L1 is also common in humans; although it is not present in as many copies as Alu, its larger size means that this element constitutes approximately 15%-17% of the human genome (Kazazian & Moran, 1998; Slotkin & Martienssen, 2007). In humans, these non-LTR TEs are the only active class of transposons; LTR retrotransposons and DNA transposons are just ancient genomic relics and are not capable of hopping.
Autonomous and non-autonomous transposons
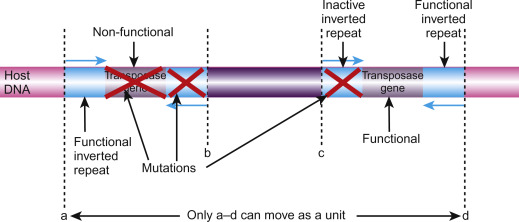
Both class 1 and class 2 TEs can be autonomous or non-autonomous. Autonomous TEs can move on their own, while non-autonomous elements require the presence of other TEs to move. This is because nonautonomous elements lack the transposase or reverse transcriptase gene needed for their transposition, so they must “borrow” these proteins from another element to move. Ac elements, for example, are autonomous because they can move on their own, while Ds elements are not autonomous because they require the presence of Ac to transpose.
What Jumping Genes Do (Besides Jumping)
The fact that about half of the human genome is made up of TEs, with a significant portion of the L1 and Alu retrotransposons, raises an important question: What do all these jumping genes do, besides jump? Much of what a transposon does depends on where it lands. Landing inside a gene can result in a mutation, as was discovered when L1 insertions into the factor VIII gene caused haemophilia (Kazazian et al., 1988). Similarly, a few years later, the researchers found L1 on the APC genes in colon cancer cells, but not on the APC genes in healthy cells in the same individuals. This confirms that L1 is transposed in mammalian somatic cells and that this element could play a causal role in disease development (Miki et al., 1992).

Silencing and Transposons
Unlike L1, most TEs appear to be silent; in other words, these elements do not produce a phenotypic effect nor do they actively move through the genome. At least that has been the general scientific consensus. Some silenced TEs are inactive because they have mutations that affect their ability to move from one chromosomal location to another; others are perfectly intact and capable of movement but are kept inactive by epigenetic defence mechanisms such as DNA methylation, chromatin remodelling, and miRNAs. In chromatin remodelling, for example, chemical modifications to chromatin proteins cause chromatin to shrink so much in certain areas of the genome that genes and TEs in those areas are silenced because transcription enzymes simply can’t access them.
Another example of transposon silencing involves plants of the genus Arabidopsis. Researchers studying these plants have discovered that they contain more than 20 different mutator transposon sequences (a type of transposon identified in maize). In wild-type plants, these sequences are methylated or silenced. However, in plants that are defective for one of the enzymes responsible for methylation, these transposons are transcribed. Furthermore, several different mutant phenotypes have been explored in methylation-deficient plants, and these phenotypes have been linked to transposon insertions (Miura et al., 2001).
Based on studies like these, scientists know that some ETs are epigenetically silenced; in recent years, however, researchers have begun to question whether certain TEs might have a role in epigenetic silencing. Interestingly, it was Barbara McClintock who first speculated that TEs might play this type of regulatory role (McClintock, 1951). Scientists have taken decades to collect enough evidence to consider that perhaps McClintock’s speculation had an ounce of truth.
Transposons can encode siRNAs that mediate their own silencing
Because transposon movement can be destructive, it is not surprising that most transposon sequences in the human genome are silent, allowing this genome to remain relatively stable, despite the prevalence of TE. In fact, the researchers believe that of the 17% of the human genome that is encoded by L1-related sequences, only about 100 active L1 elements remain. Furthermore, the research suggests that even these few remaining active transposons are inhibited from jumping in a variety of ways that go beyond epigenetic silencing.
For example, in human cells, small interfering RNAs (siRNAs), also known as RNAi, can prevent transposition. RNAi is a natural mechanism that eukaryotes often use to regulate gene expression. What is especially interesting about this situation is that the siRNAs that interfere with L1 activity are derived from the 5′ untranslated region (5′ UTR) of LTR L1. Specifically, the 5’UTR of the L1 promoter encodes a sense promoter that transcribes L1 genes, as well as an antisense promoter that transcribes antisense RNA. Yang and Kazazian (2006) showed that this results in homologous sequences that can hybridize, thus forming a double-stranded RNA molecule that can serve as a substrate for RNAi. Furthermore, when the researchers inhibited endogenous siRNA silencing mechanisms, they observed an increase in L1 transcripts, suggesting that L1 transcription is indeed inhibited by siRNA.
Transposons are not always destructive
Not all transposon jumping has harmful effects. Indeed, transposons may drive the evolution of genomes by facilitating translocation of genomic sequences, exon shuffling, and double-strand break repair. Insertions and transpositions can also alter phenotypes and gene regulatory regions. In the case of the medaka fish, for example, the DNA transposon Tol2 is directly related to pigmentation. A highly inbred line of these fish was shown to have a variety of pigmentation patterns.
In the members of this line in which the Tol2 transposon jumped “cleanly” (ie, without removing other parts of the genomic sequence), the fish were albino. But when Tol2 did not jump cleanly from the regulatory region, the result was a wide range of hereditary pigmentation patterns (Koga et al., 2006).
The fact that transposable elements are not always perfectly removed and can take up genomic sequences during the journey has also resulted in a phenomenon scientists call exon shuffling. Exon shuffling results in the juxtaposition of two previously unrelated exons, usually by rearrangement, potentially creating new gene products (Moran et al., 1999).
The ability of transposons to increase genetic diversity, coupled with the ability of the genome to inhibit most TE activity, results in a balance that makes transposable elements an important part of gene evolution and regulation in all organisms that carry these sequences.